Abstract
The project obtains Ontario Gas Demand from tsdl library and analyzed it with linear time series models and I fitted the data with SARIMA(3,0,3)(4,1,2) 12 which corrected predicted the gasoline demand of next cycle after various transformation and differencing.
1.Introduction
Given current high inflation environment and sky-rocketing gasoline price, it is of the interest to analysis how demand of Gasoline changes over time. There is a data collected in TSDL library of gasoline demand in Ontario province.
2.Data modeling
2.1 Data collection
1
2
3
4
5
library(tsdl)
options(warn=-1)
tsdl_gasdemand <- subset(tsdl, "Sales")
X_t <- tsdl_gasdemand[[1]]
attributes(X_t)
$tsp
[1] 1960.000 1975.917 12.000
$class
[1] "ts"
$source
[1] "Abraham & Ledolter (1983)"
$description
[1] "Monthly gasoline demand Ontario gallon millions 1960 – 1975"
$subject
[1] "Sales"
2.2 Data spilting
Total of 192 observations, make 12 obs as test data set and rest as train data set
1
2
3
4
5
6
train <- X_t[1:180]
test <- X_t[181:length(X_t)]
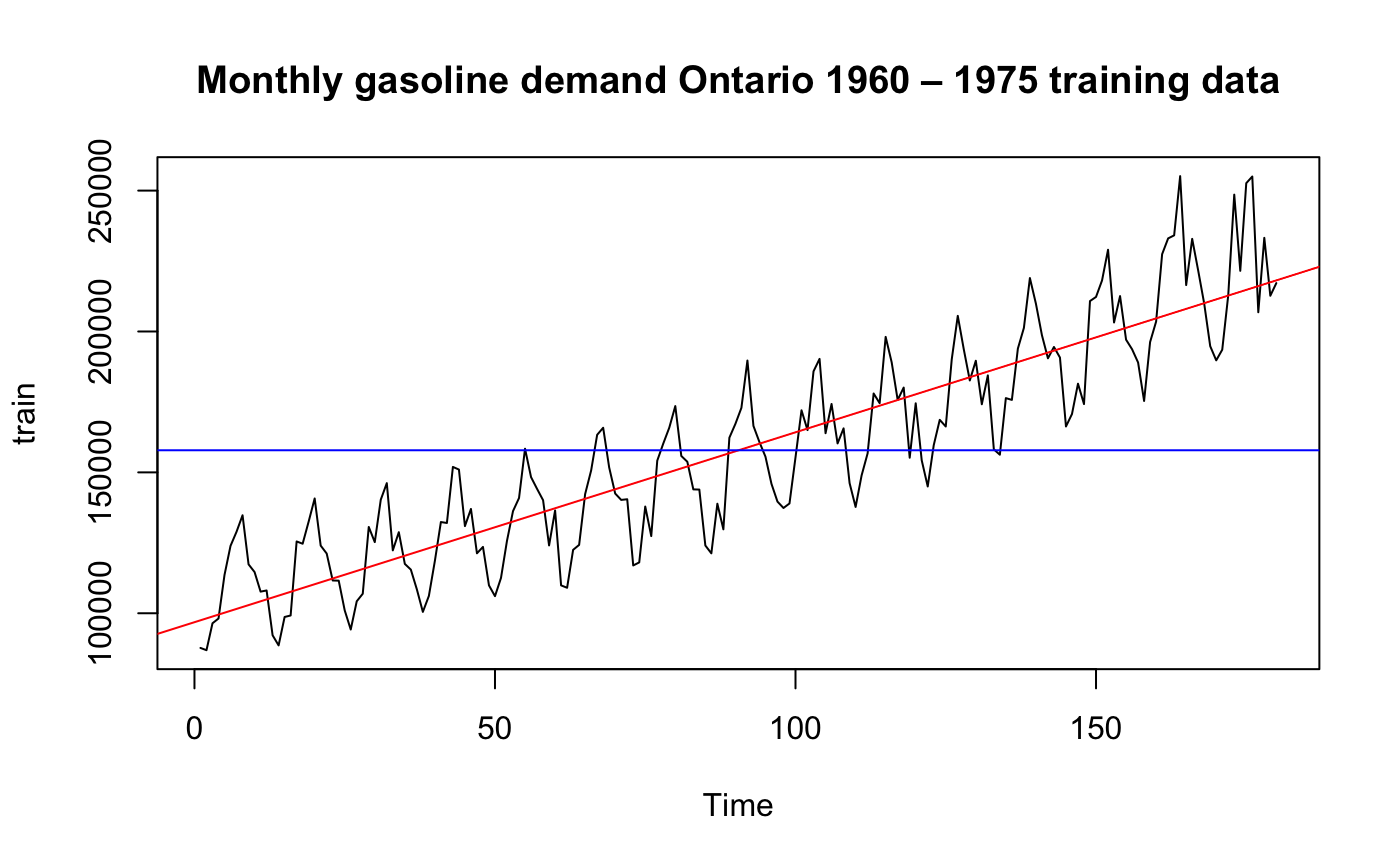
plot.ts(train, main="Monthly gasoline demand Ontario 1960 – 1975 training data")
fit <- lm(train ~ as.numeric(1:length(train)))
abline(fit, col="red")
abline(h=mean(train), col="blue")
Clearly this data has trend and it is symmetric so it may fit a linear model we have learned
2.3 Normality Check of Training data
1
2
3
4
5
6
7
8
par(mfrow=c(1,2))
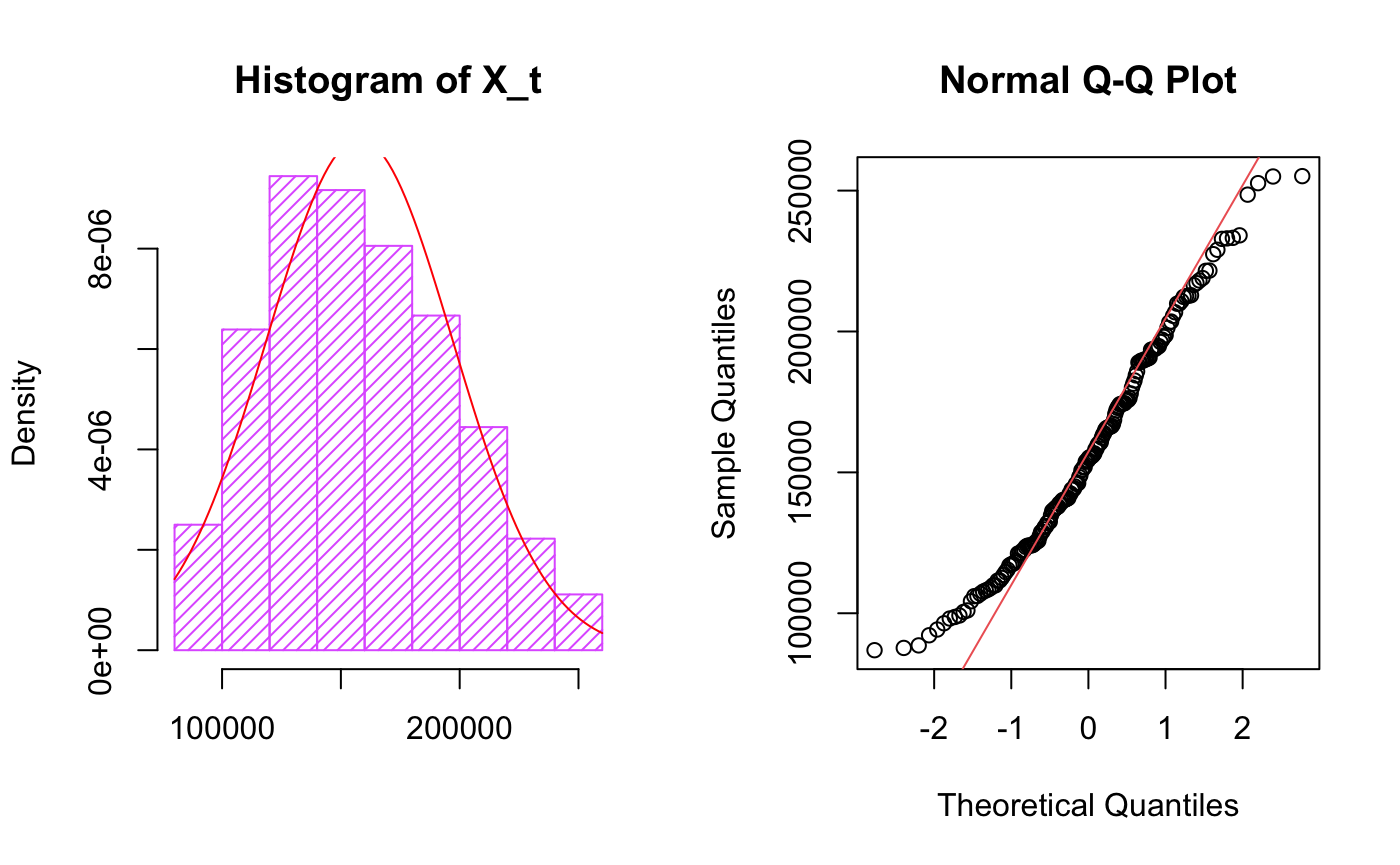
hist(train, col="mediumorchid1", xlab="",
main="Histogram of X_t",freq = F,density = 20)
m <- mean(train)
std <- sqrt(var(train))
curve(dnorm(x, m, std), col="red", add=TRUE,yaxt = "n")
qqnorm(train)
qqline(train,col="indianred2")
It follows normal distribution with some skew by looking at histogram plots, QQ plot suggests there are outliers.
2.4 ACF/PACF of Training data
1
2
3
par(mfrow=c(1,2))
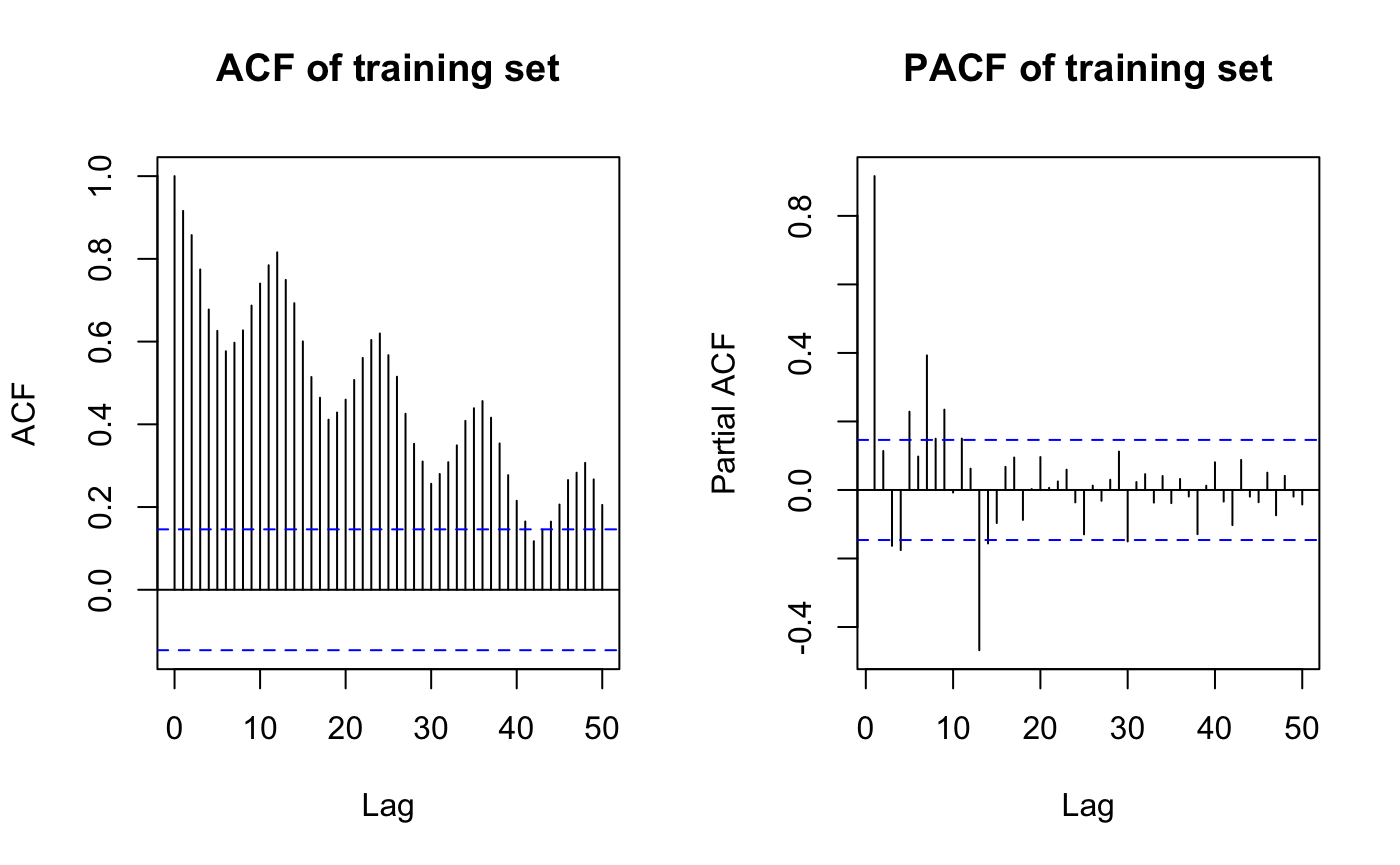
acf(train, lag.max=50, main="ACF of training set")
pacf(train, lag.max=50, main="PACF of training set")
it looks like differencing is needed as there is seasonality(monthly) in ACF and strong peak at lag12 in PACF
2.5 Decomposition and transformation of train data
1
2
3
y <- ts(as.ts(train), frequency = 12)
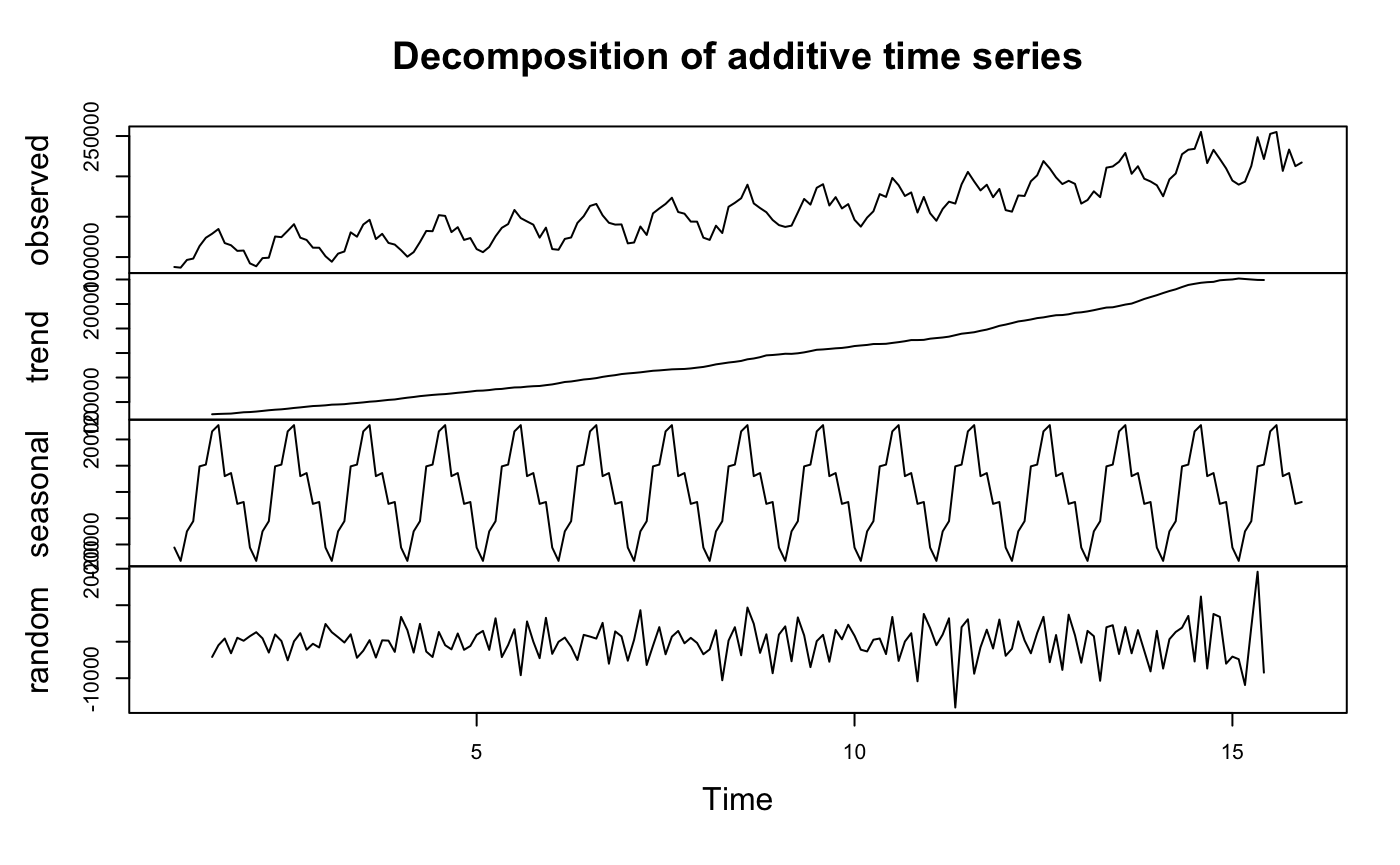
decomp <- decompose(y)
plot(decomp)
we can see there is seasonal component and trend which confirms previous conclusion again. To remove trend, transform data or take difference.
\begin{eqnarray} f_\lambda\left(U_t\right)= \begin{cases}\lambda^{-1}\left(U_t^\lambda-1\right), & U_t\geq 0, \lambda>0 \\ \ln U_t, & U_t>0, \lambda=0\end{cases} \notag \end{eqnarray}
1
2
3
4
5
6
library(MASS)
t = 1:length(train)
fit = lm(train ~ t)
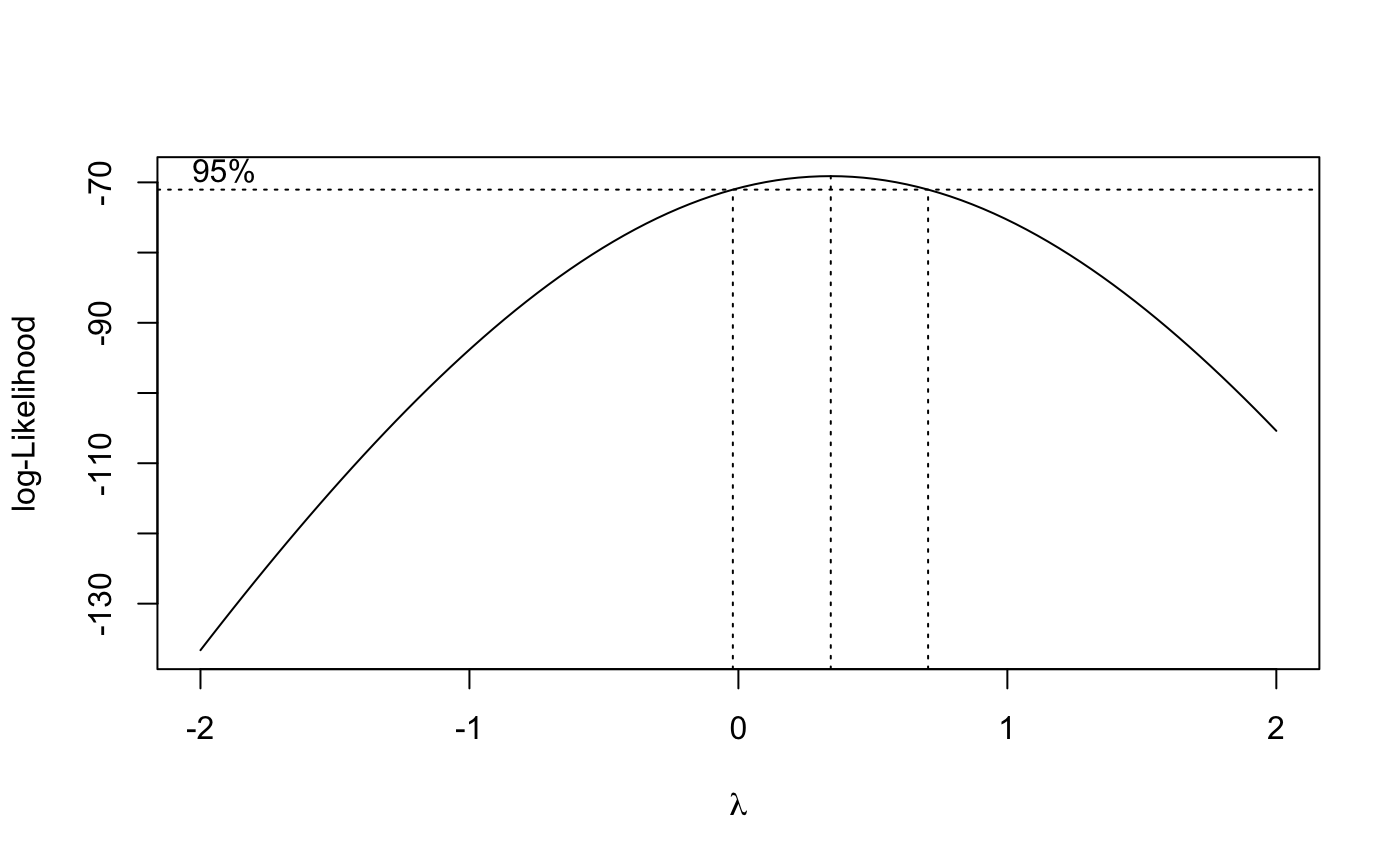
bcTransform = boxcox(train ~ t,plotit = TRUE)
lambda = bcTransform$x[which.max(bcTransform$y)]
train.log= log(train)
because box-cox test suggest log transformation($\lambda \geq 0$), I will adopt log transformation method first.
1
2
3
4
5
6
7
8
9
10
11
12
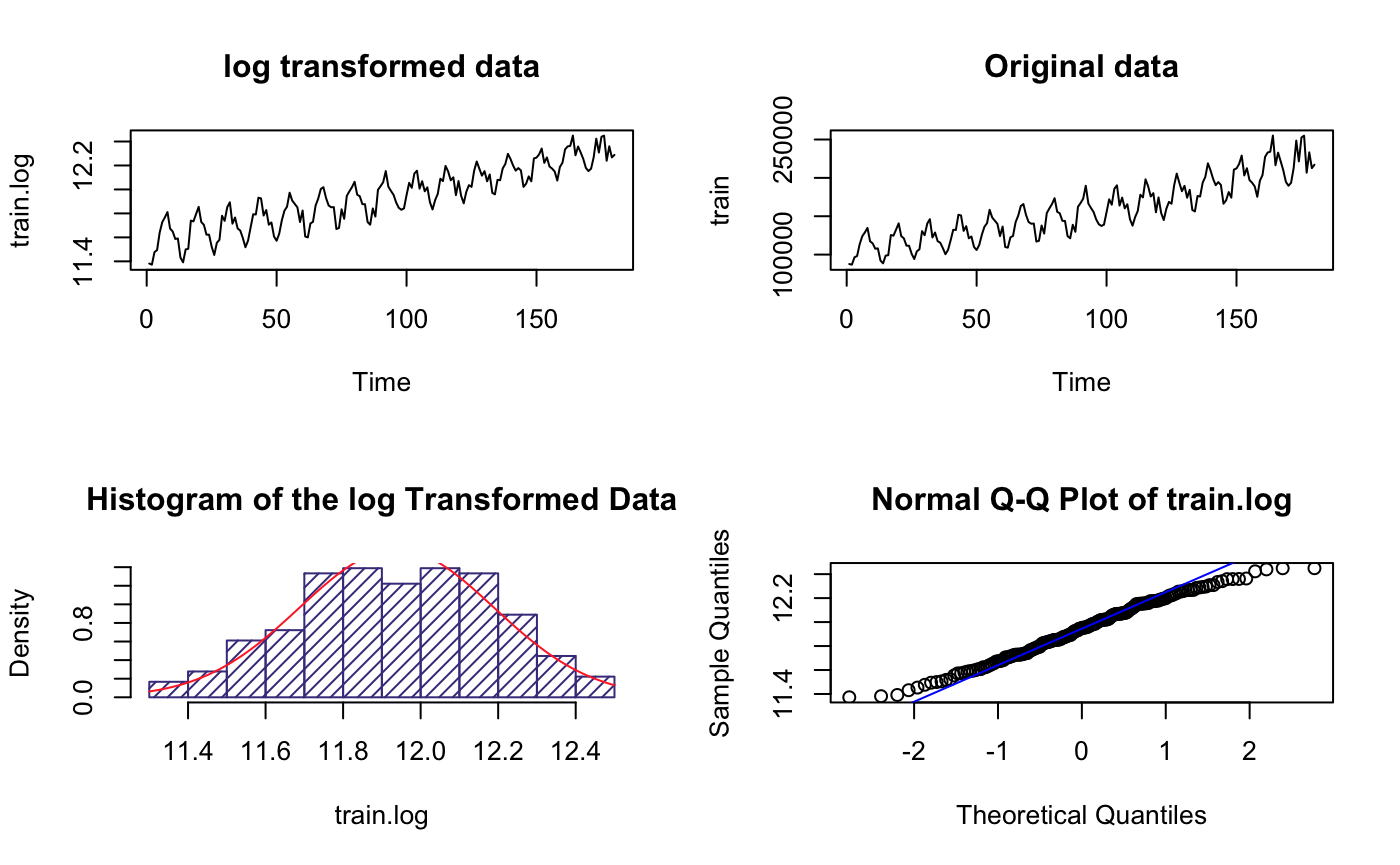
par(mfrow=c(2,2))
plot.ts(train.log,main = "log transformed data")
plot.ts(train,main = "Original data")
hist(train.log, col="slateblue4",
main="Histogram of the log Transformed Data",
freq = F,
density = 20)
m1 <- mean(train.log)
std1 <- sqrt(var(train.log))
curve(dnorm(x, m1, std1), col="firebrick1", add=TRUE)
qqnorm(train.log, main="Normal Q-Q Plot of train.log")
qqline(train.log,col ="blue")
The range(y-axis) became much smaller but others look like the same, suggesting higher variance
It’s roughly normal based on the histogram plot
1
2
3
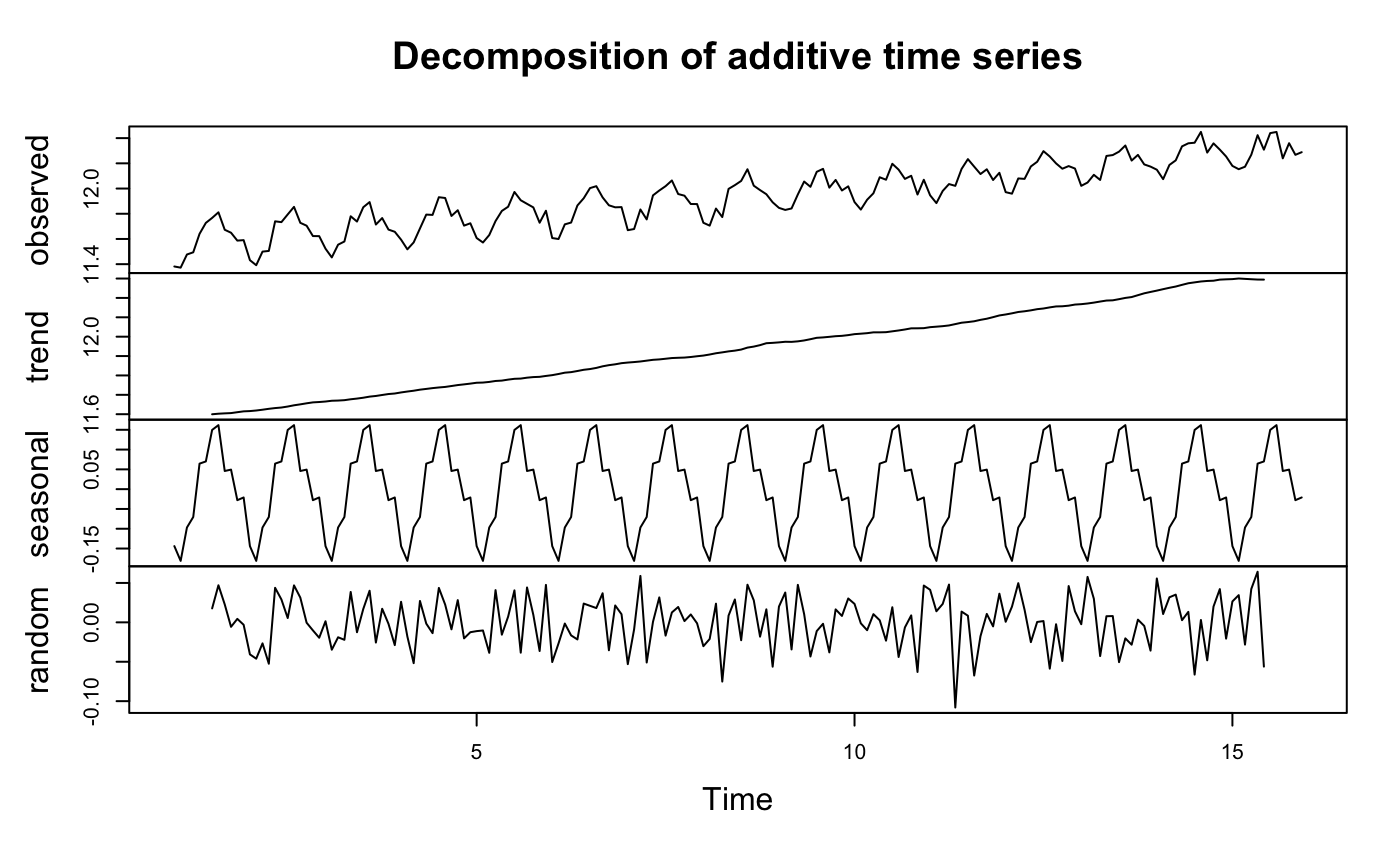
y.log <- ts(as.ts(train.log), frequency = 12)
decomp.log <- decompose(y.log)
plot(decomp.log)
It is clear we need to do difference because there is trend even after log transformation
3 Data transformation
3.1 Difference time series(log transformed)
1
2
3
4
5
6
7
8
9
10
11
12
13
14
suppressMessages(library(forecast))
diffs <- diff(train.log,12) #remove seasonality
ddiffs <- diff(diffs,1)# remove trend
ts_var <- c(var(train.log),var(diffs),var(ddiffs))
ts_var_descrb <- c("Var of log TS","Var of diffs",
"Var of ddiffs")
df <- data.frame(ts_var_descrb,ts_var)
df
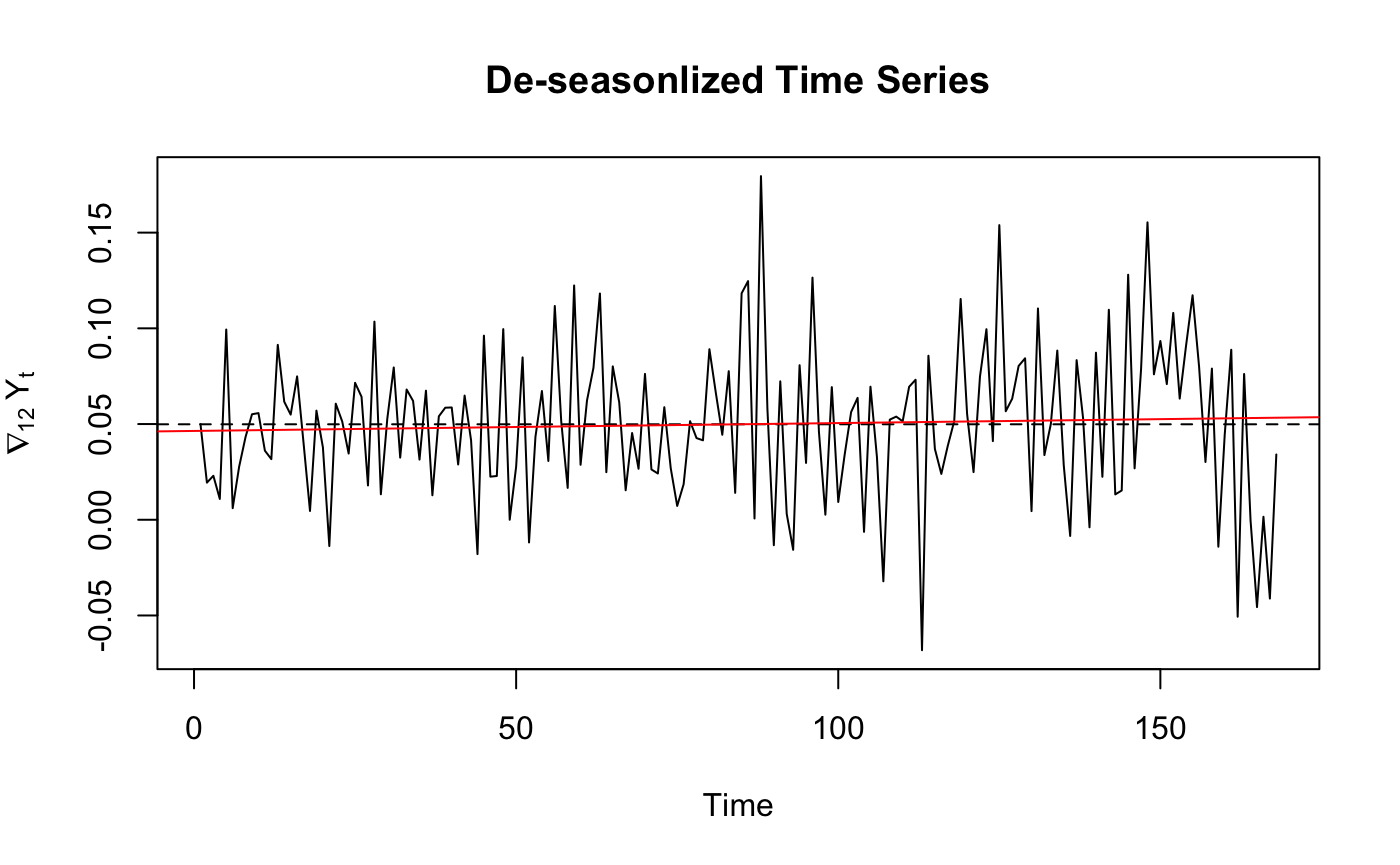
ts.plot(diffs, main="De-seasonlized Time Series",
ylab=expression(nabla[12]~Y[t]))
abline(h=mean(diffs), lty=2)
fitdiff <- lm(diffs ~ as.numeric(1:length(diffs)))
abline(fitdiff, col="red")
ts_var_descrb
<chr> ts_var <dbl>
Var of log TS 0.062973684
Var of diffs 0.001643779
Var of ddiffs 0.003746485
From the variance comparison list we can see that for log transformed Time Series , difference once at lag 12 have lowest variance.So we chose to use diffs to plot ACF/PACF.
3.2 Histagram of De-seasonal data
1
2
3
4
5
6
7
par(mfrow=c(1,2))
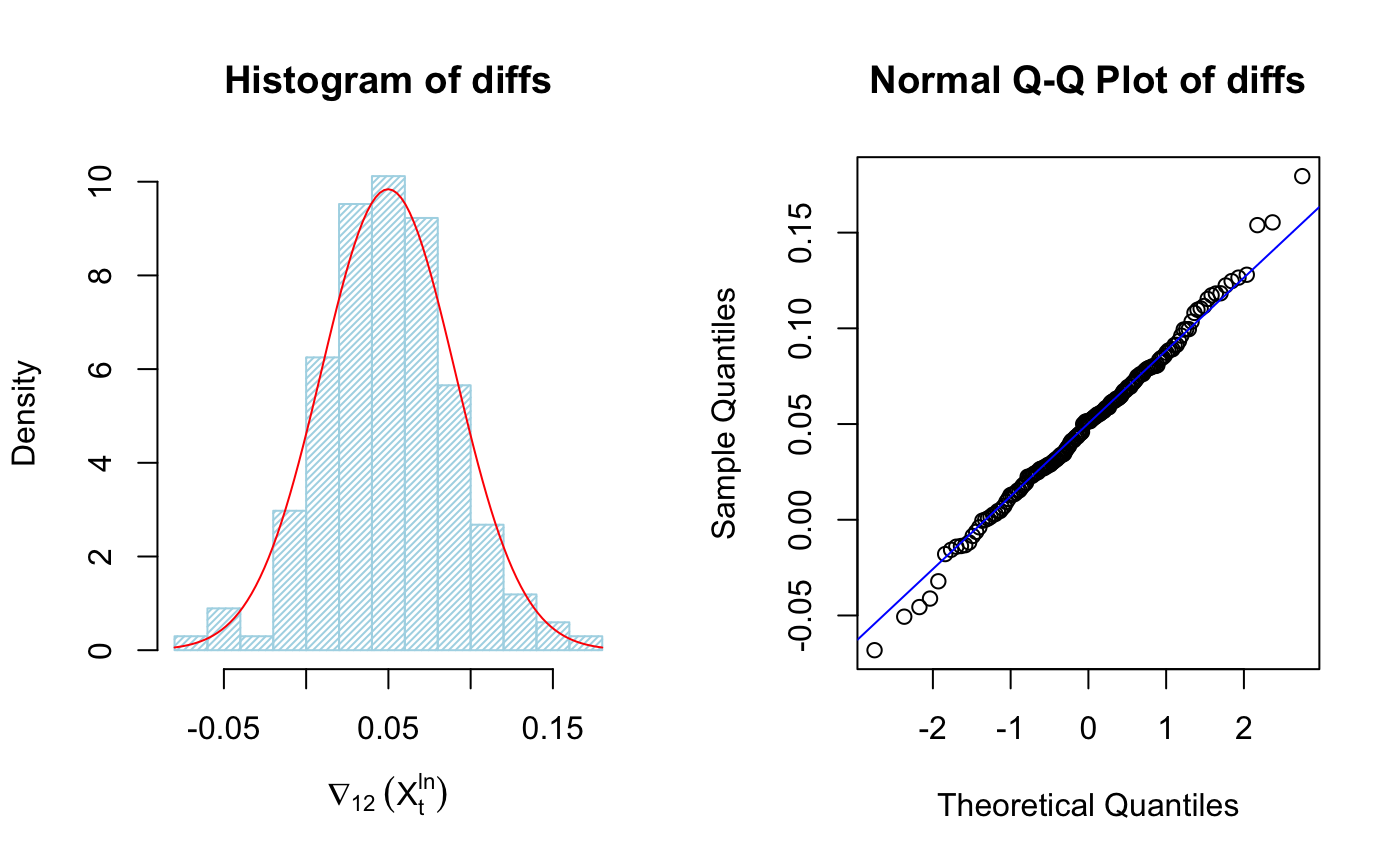
hist(diffs, col="light blue", xlab=expression(nabla[12]~(X[t]^{ln})), prob=TRUE,density = 40)
m <- mean(diffs)
std <- sqrt(var(diffs))
curve(dnorm(x,m,std), add=TRUE,col="red")
qqnorm(diffs, main="Normal Q-Q Plot of diffs")
qqline(diffs,col ="blue")
It’s clear the data is much more “Gaussian”, we can plot ACF/PACF of differenced transformed data
3.3 ACF/PACF of differenced trainging data
1
2
3
par(mfrow=c(1,2))
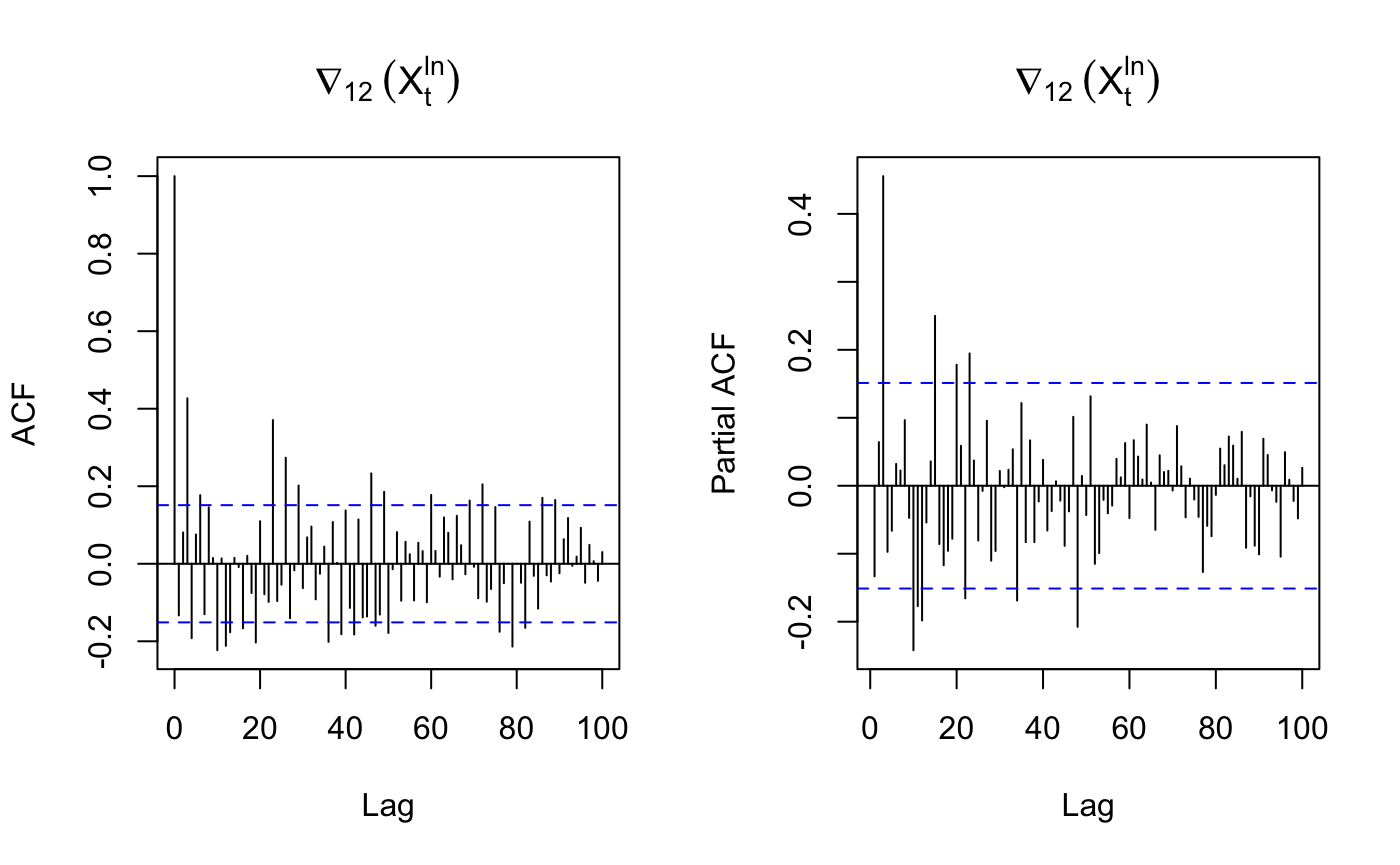
acf(diffs, lag.max=100, main=expression(nabla[12]~(X[t]^{ln})))
pacf(diffs, lag.max=100, main=expression(nabla[12]~(X[t]^{ln})))
3.4 Model parameter identification
seasonal part
D=1 – Differenced once at lag 12
s= 12 – 12 month per year
P=4 – strong peak at multiple of lag 12 before lag 60 in PACF
Q could be 1,2,3,4 based on the multiple of ACF.
non-seasonal part
p = 3 – strong peak at lag 3
d = 0 – not differenced at lag 1
q = 3 – strong peak at lag 3 in ACF
That gives SARIMA(3,0,3)x(4,1,c(1,2,3,4))12
4.0 Model fitting
1
2
3
4
5
6
suppressMessages(library(astsa))
model1 <- sarima(xdata=train.log, p=3, d=0, q=3, P=4, D=1, Q=1, S=12, details = F)
model2 <- sarima(xdata=train.log, p=3, d=0, q=3, P=4, D=1, Q=2, S=12, details = F)
model3 <- sarima(xdata=train.log, p=3, d=0, q=3, P=4, D=1, Q=3, S=12, details = F)
model4 <- sarima(xdata=train.log, p=3, d=0, q=3, P=4, D=1, Q=4, S=12, details = F)
4.1 AICC check
1
2
3
4
AICc_value <- c(model1$AICc,model2$AICc,model3$AICc,model4$AICc)
AICc_des <- c("AICC of Model1","AICC of Model2","AICC of Model3","AICC of Model4")
df_AICC <- data.frame(AICc_des,AICc_value)
df_AICC
1
2
3
4
5
6
AICc_des AICc_value
<chr> <dbl>
AICC of Model1 -4.076983
AICC of Model2 -4.135593
AICC of Model3 -4.133020
AICC of Model4 -4.129460
Based on AICC value, we can chose model 2 which is SARIMA(3,0,3)(4,1,2)$_{12}$
1
2
model2$fit
model2$AICc
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
Call:
arima(x = xdata, order = c(p, d, q), seasonal = list(order = c(P, D, Q), period = S),
xreg = constant, transform.pars = trans, fixed = fixed, optim.control = list(trace = trc,
REPORT = 1, reltol = tol))
Coefficients:
ar1 ar2 ar3 ma1 ma2 ma3 sar1 sar2 sar3 sar4 sma1
-0.2531 -0.1308 0.2677 0.2831 0.3681 0.2317 0.1603 -0.6132 -0.4573 -0.3213 -0.9767
s.e. 0.1585 0.1437 0.1620 0.1655 0.1380 0.1685 0.2127 0.0834 0.0829 0.1392 0.4181
sma2 constant
0.9043 0.0043
s.e. 0.5322 0.0001
sigma^2 estimated as 0.0005733: log likelihood = 362.57, aic = -697.14
[1] -4.135593
We need to modify the model because some Coefficient have 0 in their confidence interval.
1
2
3
4
model2_modi <- sarima(xdata=train.log, p=3, d=0, q=3, P=4, D=1, Q=2, S=12,
fixed = c(0,0,0,0,NA,NA,NA,NA,NA,NA,0,0,NA), details = F)
model2_modi$fit
model2_modi$AICc
1
2
3
4
5
6
7
8
9
10
11
12
Call:
arima(x = xdata, order = c(p, d, q), seasonal = list(order = c(P, D, Q), period = S),
xreg = constant, transform.pars = trans, fixed = fixed, optim.control = list(trace = trc,
REPORT = 1, reltol = tol))
Coefficients:
ar1 ar2 ar3 ma1 ma2 ma3 sar1 sar2 sar3 sar4 sma1 sma2 constant
0 0 0 0 0.2555 0.4669 -0.5696 -0.4210 -0.5124 -0.5000 0 0 0.0042
s.e. 0 0 0 0 0.0752 0.0664 0.0711 0.0796 0.0743 0.0734 0 0 0.0001
sigma^2 estimated as 0.0007702: log likelihood = 353.68, aic = -691.36
[1] -4.111059
AR1, AR2, AR3, MA1, SMA1, SMA2 term all have 0 in their CI so I have modified them to 0.
The AICC of modified model is higher than original model so we need to use the original model 2 because the coefficients are significant if the AICC of the original model is lower.
4.2 Final Model
so the model selection is
\begin{equation} SARIMA(3,0,3)(4,1,2)_{12} \notag \end{equation}
\begin{equation} (1+0.2531B^1+0.1308B^2-0.2677B^3)(1-0.1603B^{12}+0.6132B^{24}+0.4573B^{36}+0.3213B^{48})X_t=\ \notag \end{equation}
\begin{equation} (1+0.2831B+0.3681B^2+0.2317B^3)(1-0.9767B^{12}+0.9043B^{24})Z_t \notag \end{equation}
5.0 Diagnostic Checking
5.1 Unit Root
1
2
3
4
5
6
library(UnitCircle)
par(mfrow=c(2,2))
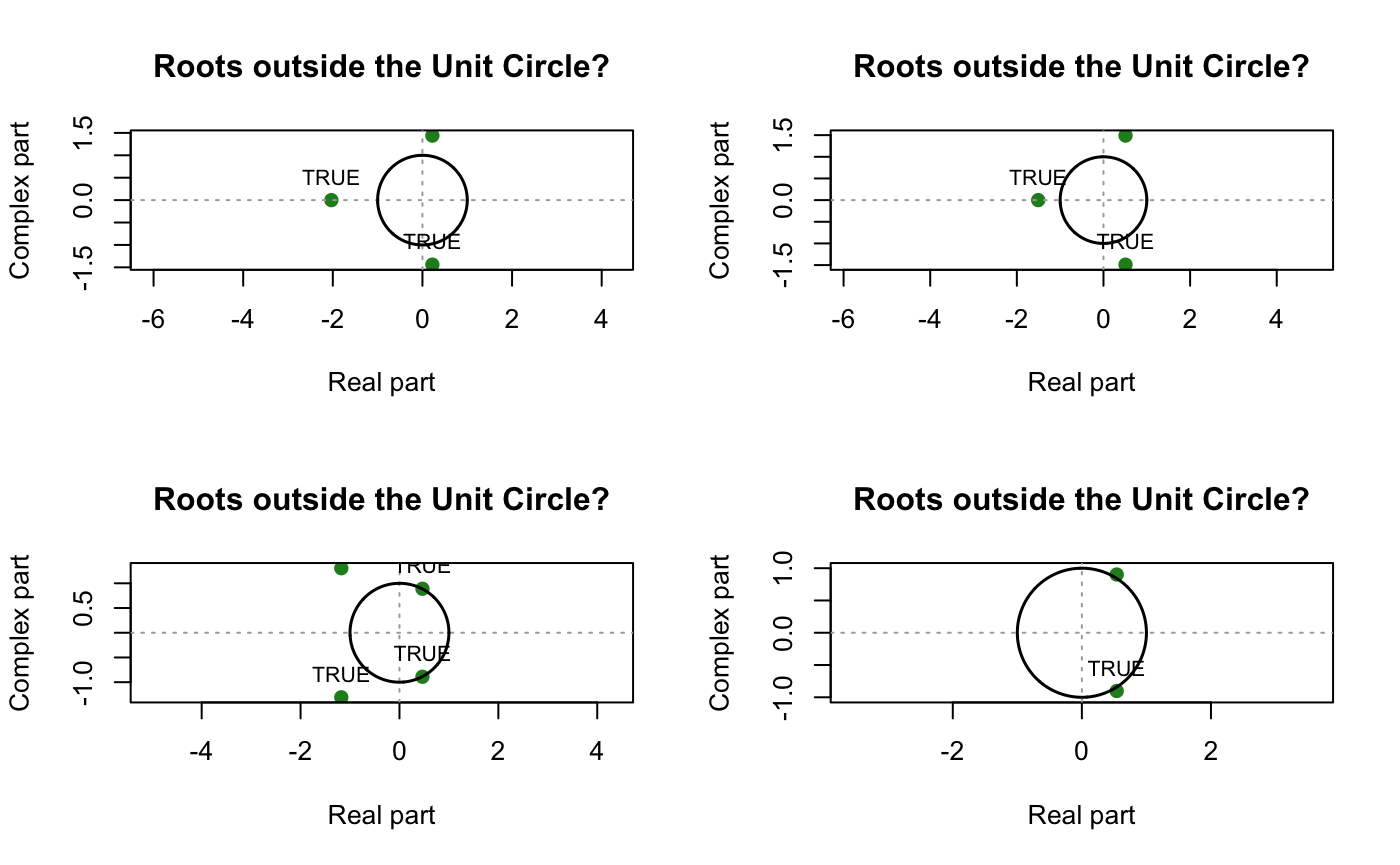
uc.check(pol_ = c(1,0.2831,0.3681,0.2317), plot_output = TRUE,print_output = F) # Non-Seasonal MA Part
uc.check(pol_ = c(1,0.2531,0.1308,0.2677), plot_output = TRUE,print_output = F)# non-seasonal AR part
uc.check(pol_ = c(1,-0.1603,0.6132,0.4573,0.3213), plot_output = TRUE,print_output = F)# seasonal AR part
uc.check(pol_ = c(1,-0.9767,0.9043), plot_output = TRUE,print_output = F)# seasonal MA part
Neither parts has unit root, we proceed to Sanity checking
5.2 Residual Normality checking
1
2
3
4
5
6
7
res <- residuals(model2$fit)
res <- tsclean(res)
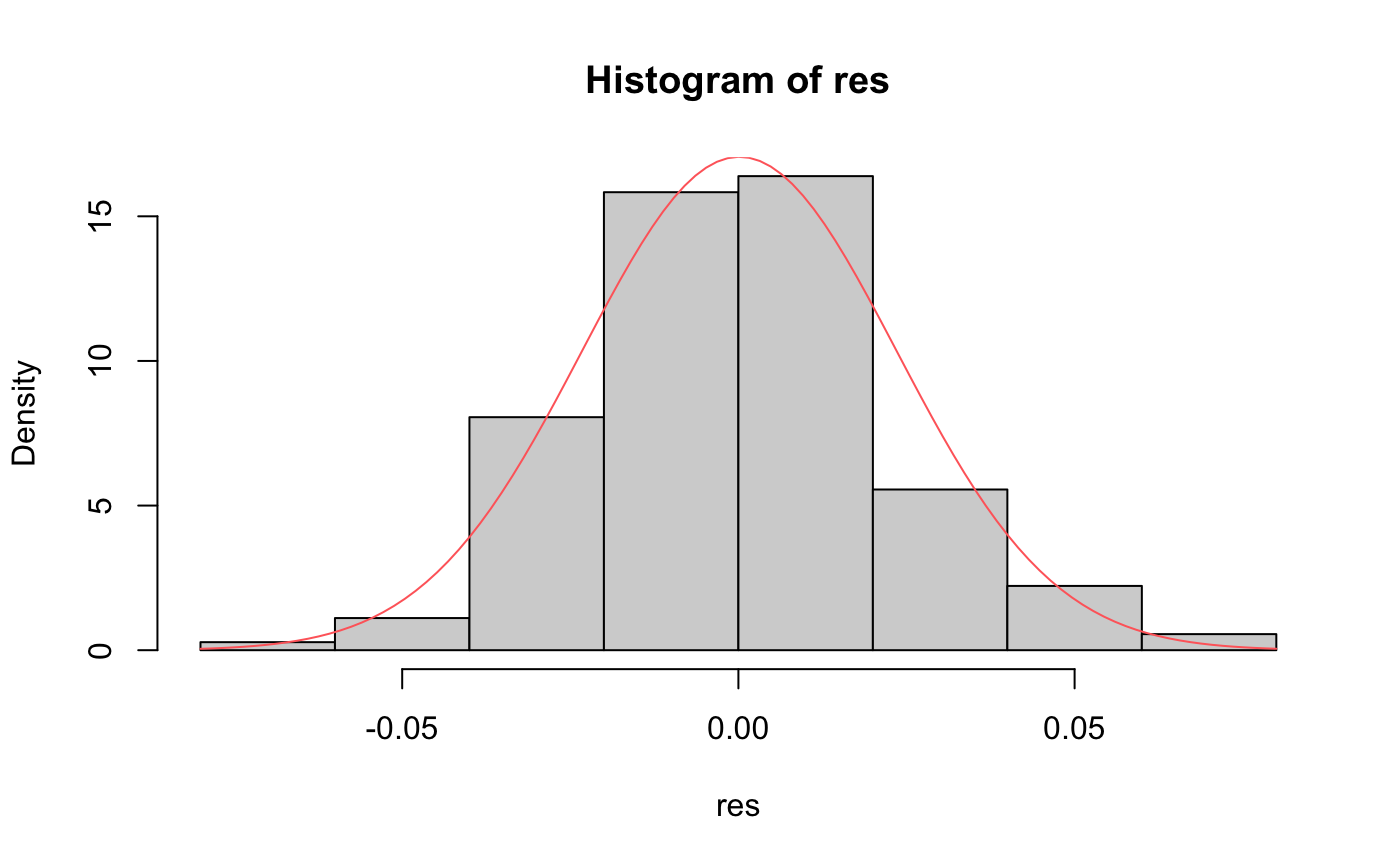
hist(res,freq = F)
m_res <- mean(res)
std_res <- sqrt(var(res))
curve(dnorm(x,m_res,std_res), add=TRUE,col="indianred1")
shapiro.test(res)
1
2
3
Shapiro-Wilk normality test
data: res
W = 0.98834, p-value = 0.1452
Shapiro-Wilk normality test
\begin{eqnarray} \begin{cases} H_0: \text{Residuals are normally distributed} \\H_1: \text{not } H_0\end{cases} \end{eqnarray}
it looks like our residuals are normally distributed based on P-value of Shaprio test (>0.05)
1
2
3
4
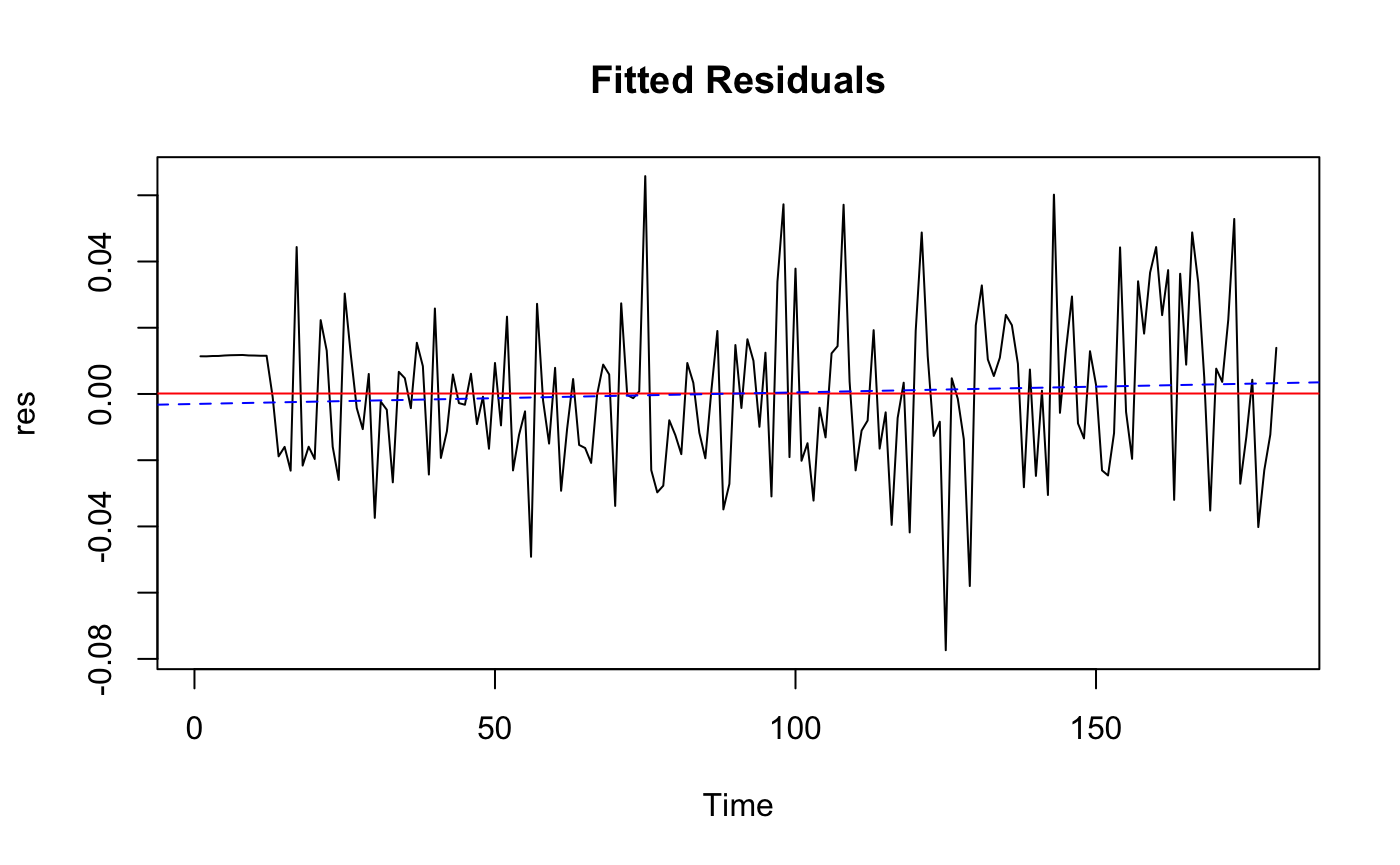
ts.plot(res, main = "Fitted Residuals")
abline(h=mean(res), col = "red")
fitres <- lm(res ~ as.numeric(1:length(res)))
abline(fitres, col="blue", lty=2)
The residual is Roughly WN
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
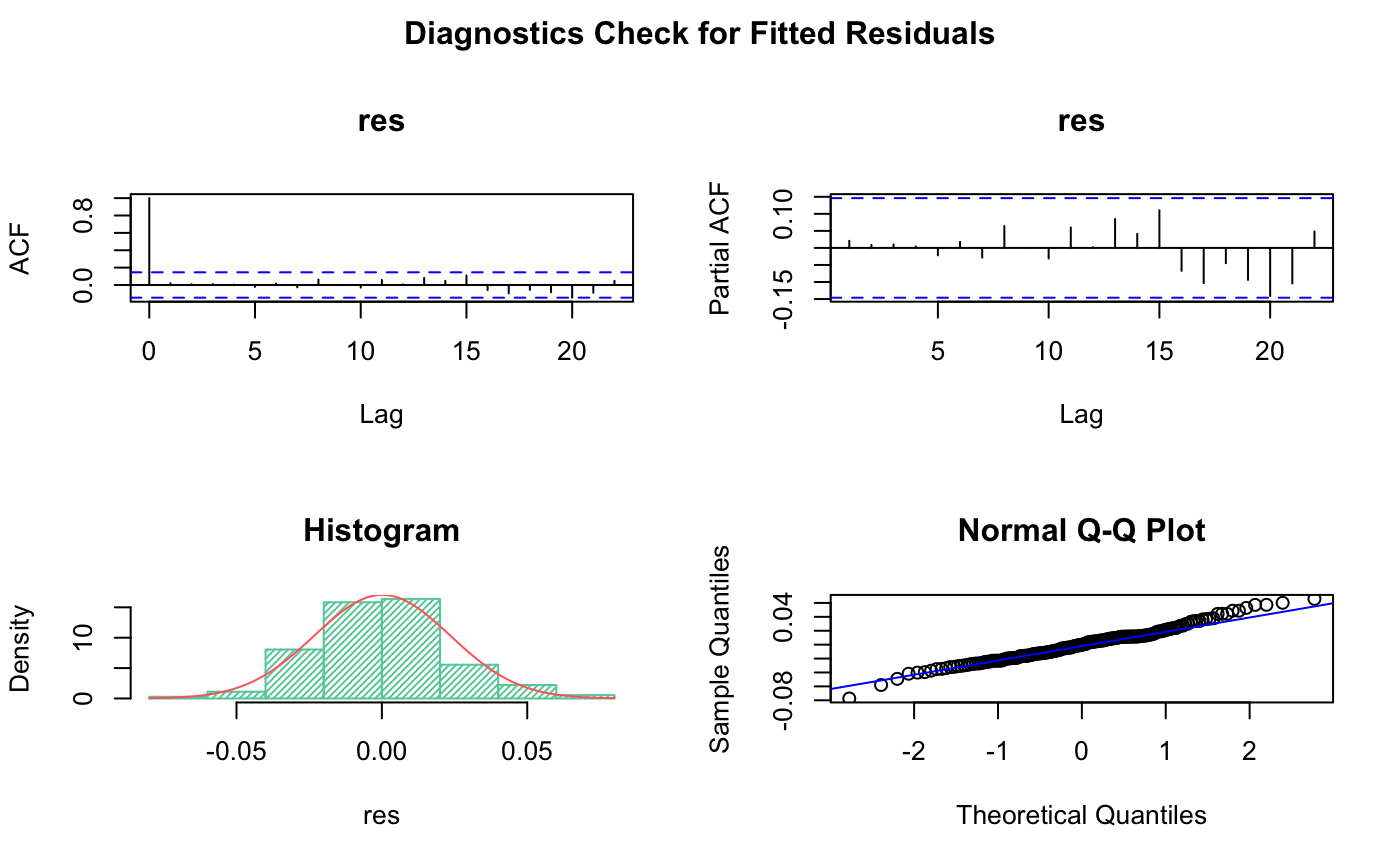
# Plot diagnostics of residuals
par(oma=c(0,0,2,0))
par(mfrow=c(2,2))
acf(res,main = "res")
pacf(res,main = "res")
# Histogram
hist(res,main = "Histogram",freq = F,density = 40,col = "aquamarine3")
m_res <- mean(res)
std_res <- sqrt(var(res))
curve(dnorm(x,m_res,std_res), add=TRUE,col="indianred1")
# q-q plot
qqnorm(res)
qqline(res,col ="blue")
# Add overall title
title("Diagnostics Check for Fitted Residuals", outer=TRUE)
From Histogram and QQ plot we can see the residual is roughly normally distributed.
5.2 Various Box Test
since there are 6 effective coefficient so we take fitdf = 6
1
2
3
Box.test(res, lag = 13, type = "Box-Pierce", fitdf = 12) #sqrt(180) is roughly 13
Box.test(res, lag = 13, type = "Ljung-Box", fitdf = 12)
Box.test((res)^2, lag = 12, type = "Ljung-Box", fitdf = 0)#McLeod-Li
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
Box-Pierce test
data: res
X-squared = 3.0571, df = 1, p-value = 0.08039
Box-Ljung test
data: res
X-squared = 3.2782, df = 1, p-value = 0.0702
Box-Ljung test
data: (res)^2
X-squared = 8.8323, df = 12, p-value = 0.7172
Because the observation of training set is 180, its square root is about 13, so I set lag = 13 in the box tests. Based on three test p-values(>0.05), we can conclude that we fail to reject the null hypothesis thus the Residual is uncorrelated of each other.
6.0 Spectral Analysis
6.1 Periodogram & Kolmogorov-Smirnov Test
1
2
3
par(mfrow=c(1,2))
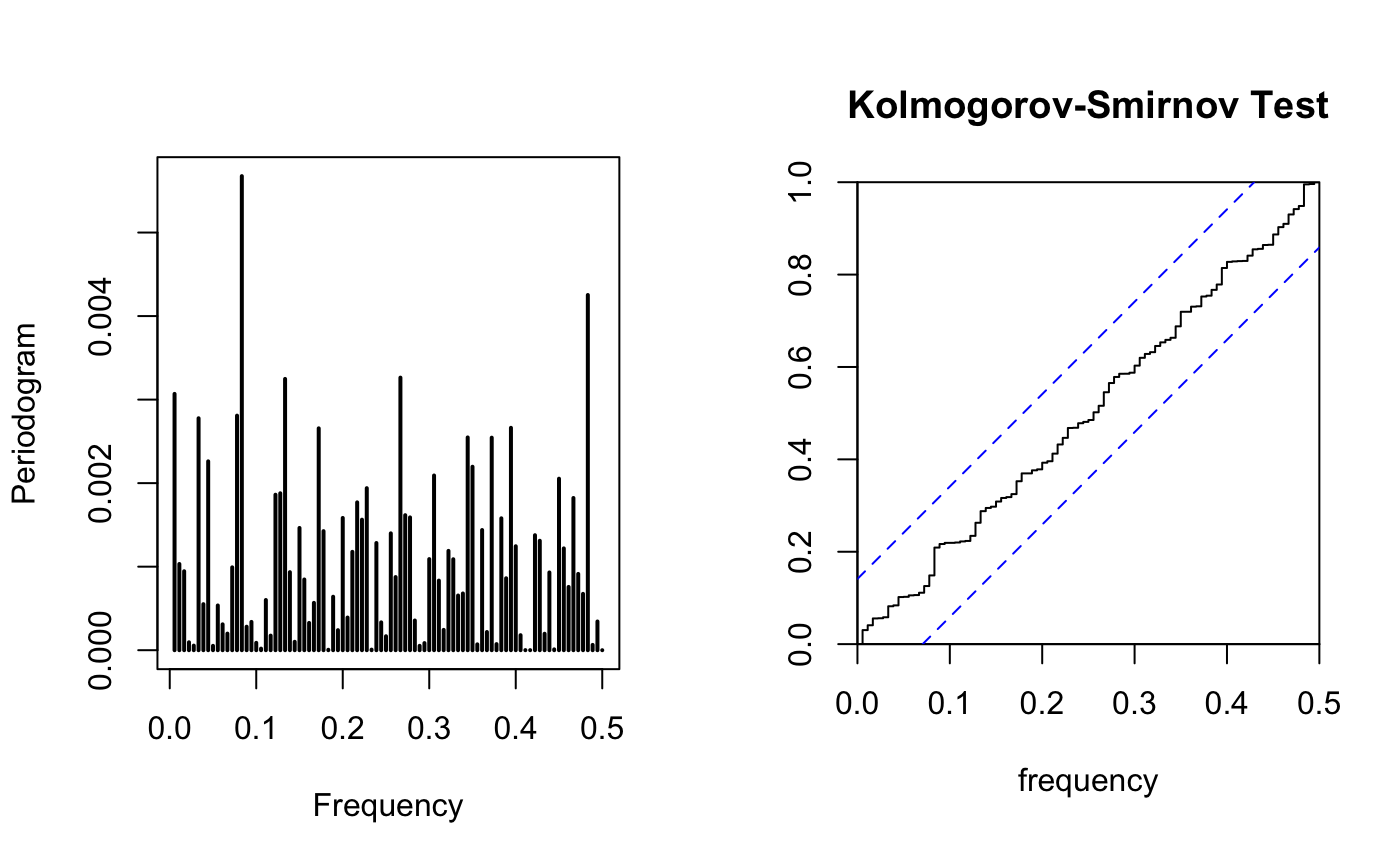
TSA::periodogram(res,plot = T)
cpgram(res,main="Kolmogorov-Smirnov Test")
all residual fall between the CI, the residual is Gaussian WN. From the periodogram, I don’t see a dominated frequency
6.2 Fisher test
1
2
suppressMessages(library(GeneCycle))
fisher.g.test(res)
1
[1] 0.3939237
since the p-value is larger than 0.05 ,original data is Gaussian distributed(Do not reject $H_0$)
7.0 Forecasting
7.1 set Confidence interval
1
2
3
4
5
SARIMA <- arima(train.log, order=c(3,0,3), method="ML",
seasonal = list(order = c(4,1,2), period = 12))
pred.demand <- predict(SARIMA, n.ahead = 12)
Upperbound <- pred.demand$pred + 1.96*pred.demand$se
Lowerbound <- pred.demand$pred - 1.96*pred.demand$se
7.2 Forecast for training data
1
2
3
4
5
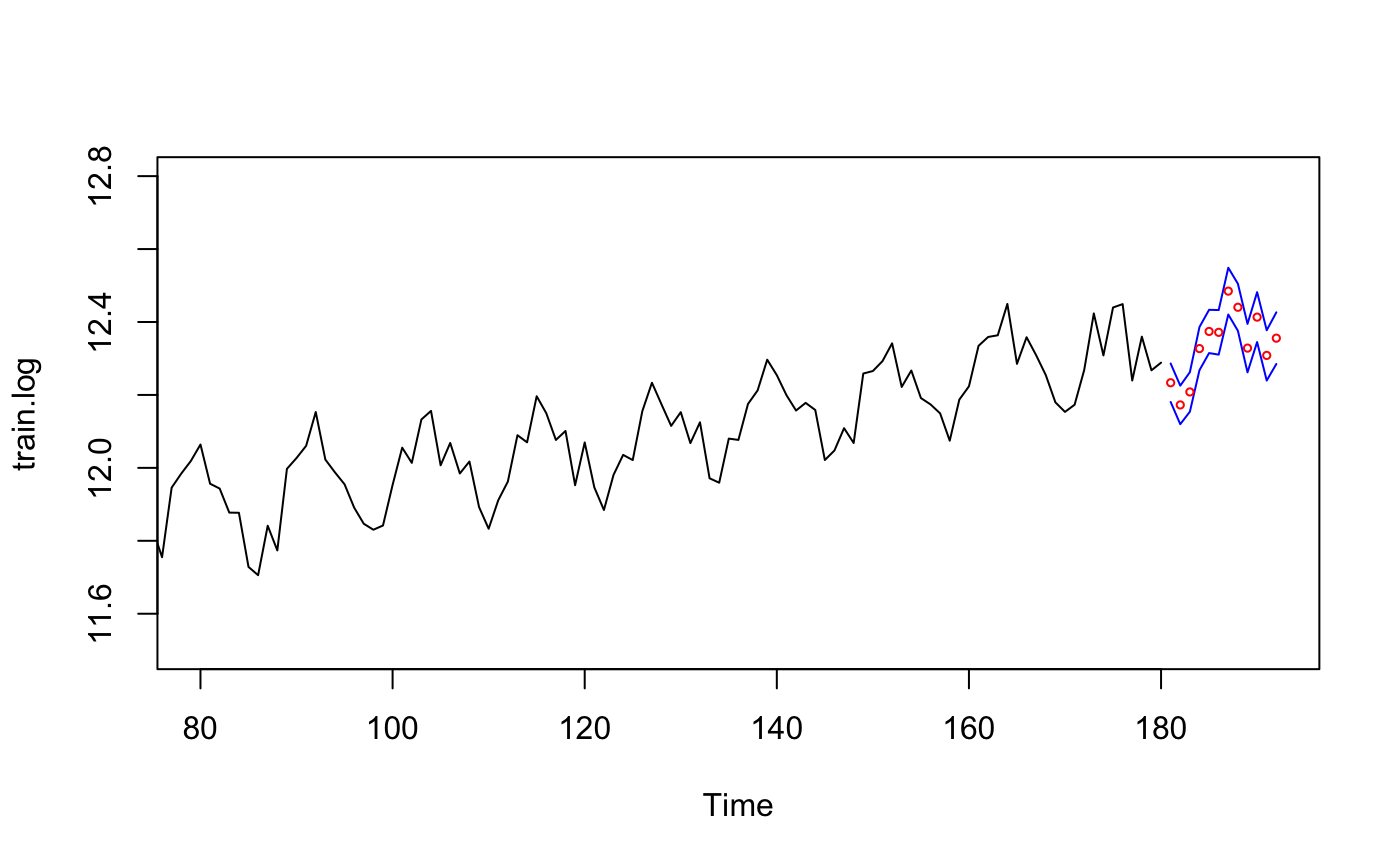
ts.plot(train.log, xlim=c(80,length(train.log)+12), ylim = c(11.5,12.8))
lines(Upperbound, col="blue")
lines(Lowerbound, col="blue")
points((length(train.log)+1):(length(train.log)+12),
pred.demand$pred, col="red",cex=0.5) # Predicted Value
7.3 Forecasted v.s true value
1
2
3
4
5
6
7
8
9
10
11
pred.orig <- exp(pred.demand$pred) # transform back
U <- exp(Upperbound)
L <- exp(Lowerbound)
ts.plot(train, xlim = c(150,length(train)+12), ylim = c(min(train),max(U)), col="black")
lines(U, col="grey", lty="dashed")
lines(L, col="grey", lty="dashed")
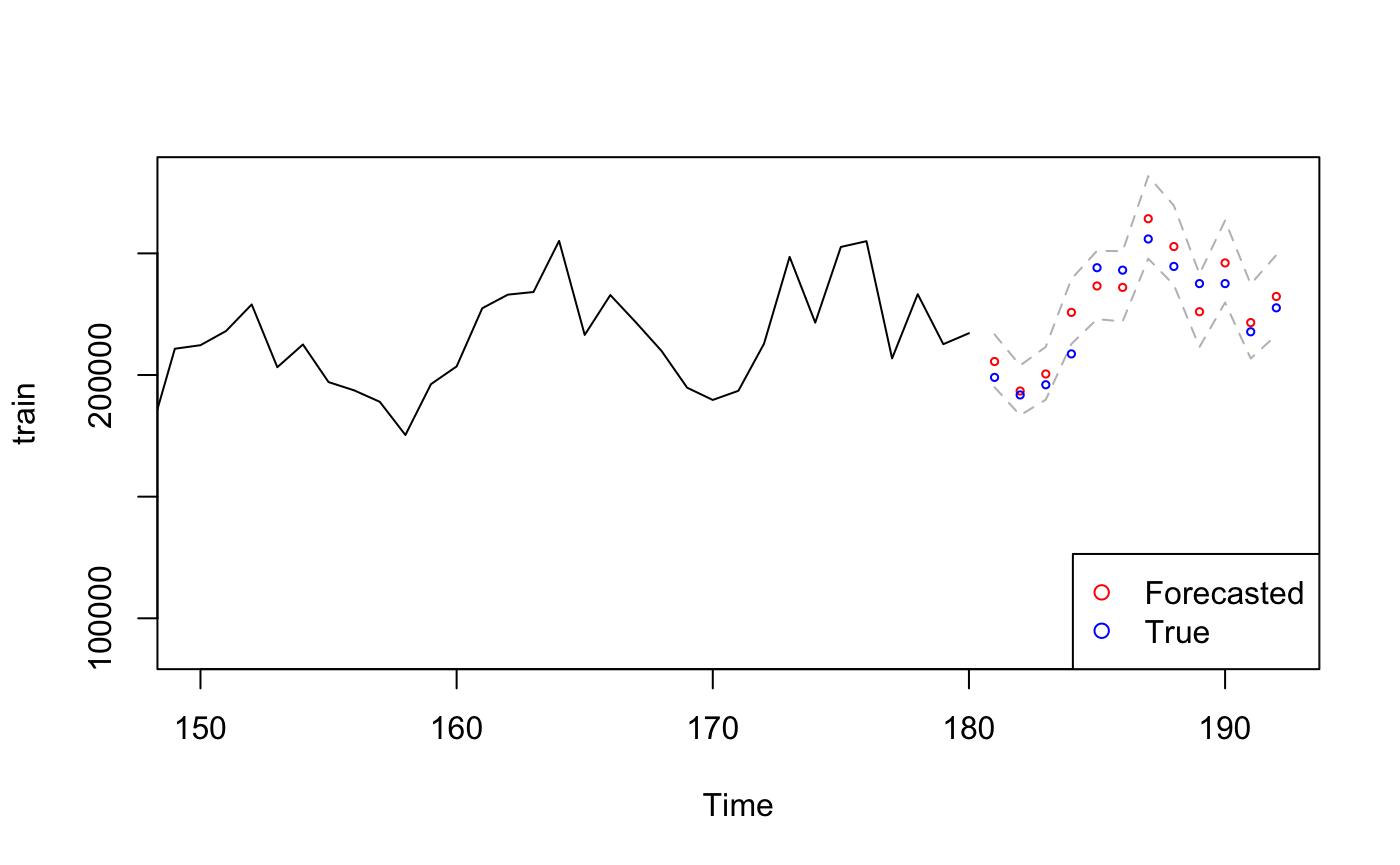
points((length(train)+1):(length(train)+12), pred.orig, col="red", cex=0.5) # Forecasted Value
points((length(train)+1):(length(train)+12), test, col="blue", cex=0.5) # True Value
# legend("bottomleft",pch = 1,col = c("red","blue"))
PerformanceAnalytics::legend("bottomright", pch=1, col = c("red", "blue"),
legend = c("Forecasted","True"))
The true value falls in 95% CI of the SARIMA model which indicating the prediction is accurate, the first predicted value even overlaps with the true value which suggesting our model is very good.
8.0 Conclusion
The purpose of this project is predicting future gas demand based on known data sequence, Throughout the process of transforming, differencing, model selection, diagnostic checking, and spectral analysis, the final model I picked is $ SARIMA(3,0,3) \times (4,1,2)_{12} $
Reference
Lecture slides & notes
https://stat.ethz.ch/R-manual/R-devel/library/stats/html/box.test.html
https://cran.r-project.org/web/packages/forecast/forecast.pdf
https://cran.r-project.org/web/packages/TSA/TSA.pdf
https://stat.ethz.ch/R-manual/R-devel/library/stats/html/arima.html
https://github.com/nickpoison/astsa/blob/master/fun_with_astsa/fun_with_astsa.md#arima-simulation
https://cran.r-project.org/web/packages/astsa/astsa.pdf